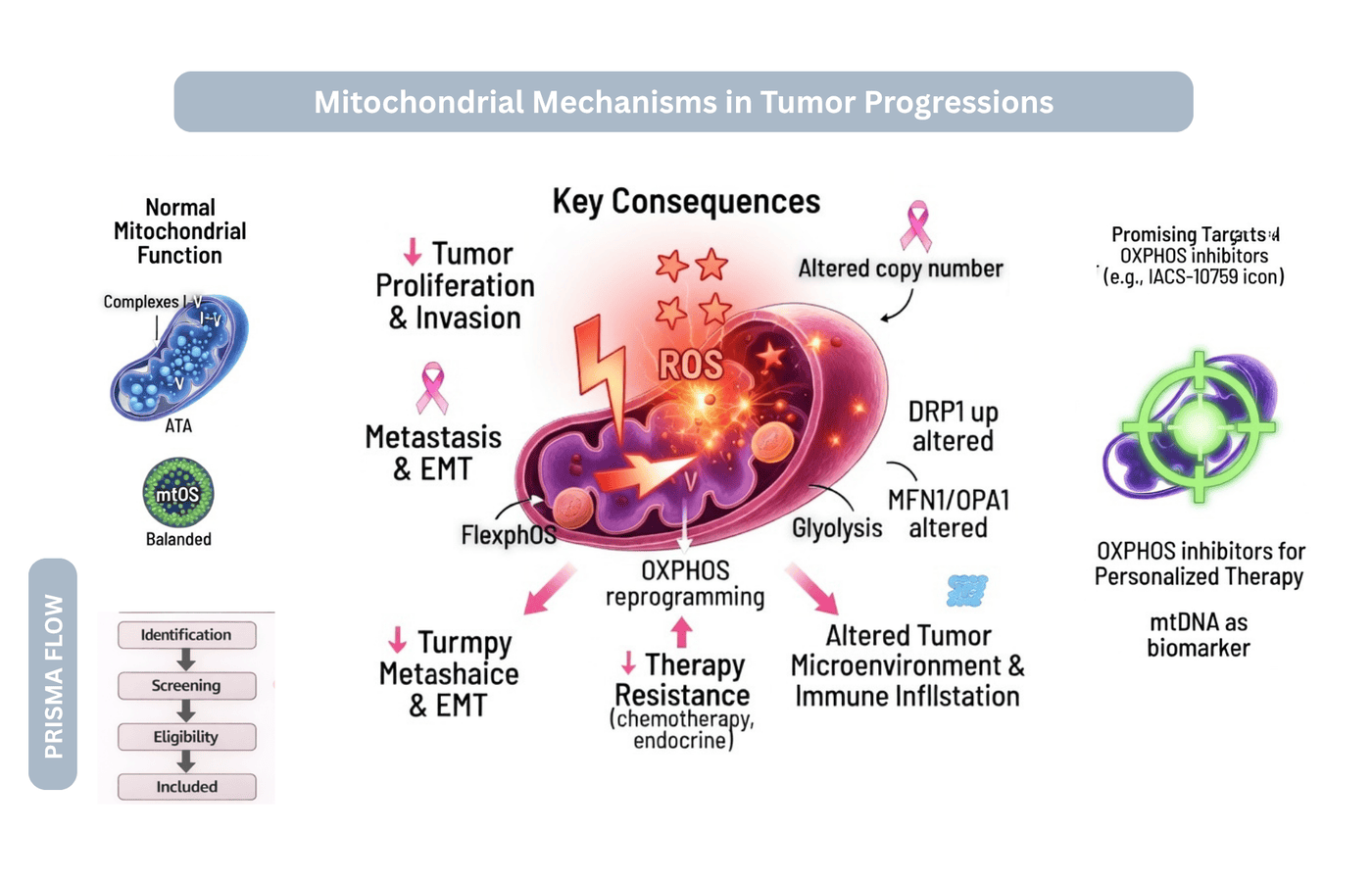
Mitochondrial DNA Mutations and Oxidative Phosphorylation Reprogramming in Colorectal Cancer and Breast Cancer: Systematic Review of Distinct Mitochondrial Mechanisms in Tumor Progressions
Keywords:
Mitochondrial Dysfunction, Oxidative Phosphorylation, Mitochondrial DNA, Reactive Oxygen Species, Breast Cancer, Colorectal Cancer, NeoplasmsAbstract
Background: Mitochondrial dysfunction with mitochondrial DNA (mtDNA) mutations and oxidative phosphorylation (OXPHOS) reprogramming have a significant role in cancer progression. The aim of the study was to mechanically compare different mechanisms of mitochondria in tumour development, both in breast and colorectal cancer. Methodology: It was a systematic review based on PRISMA guidelines 2020. The databases used in the search were PubMed, Scopus, Web of Science, and Google Scholar from 2020-2026. This was done by including clinical, experimental in vitro and in vivo studies that examined the role of mitochondrial DNA mutations and OXPHOS pathways in breast and colorectal cancer and excluding reviews, case reports, non-English studies, and irrelevant disease models. Newcastle-Ottawa Scale (NOS), Joanna Briggs Institute (JBI) checklist, ROBINS-I tool, modified in vitro tools, and SYRCLE tool were all used to assess risk of bias, with some certainty of evidence assessed using the GRADE framework. Results: 12 studies meet the inclusion criteria. Results have shown that OXPHOS reprogramming and the presence of the mutations in the mitochondrial DNA resulted in electron transport dysfunction, reactive oxygen species generation, and ATP production, all of which were contributing factors to tumour progression, metabolic adaptation, and therapeutic resistance. Mitochondrial pathways also had an effect on tumor microenvironment and prognosis. The risk of bias was moderate in general and the certainty of evidence was low according to GRADE assessment. Conclusion: Mitochondrial changes are the main focus of cancer development and are potential therapeutic targets. It is recommended that future research should be done on clinical validation and development of mitochondrial-targeted therapy to treat cancer on a personal basis.
Downloads
References
1. Zhang L, Zhang W, Li Z, Lin S, Zheng T, Hao B, et al. Mitochondria dysfunction in CD8+ T cells as an important contributing factor for cancer development and a potential target for cancer treatment: a review. J Exp Clin Cancer Res. 2022;41(1):227. https://doi.org/10.1186/s13046-022-02439-6
2. Vaupel P, Multhoff G. Revisiting the Warburg effect: historical dogma versus current understanding. J Physiol. 2021;599(6):1745–1757. https://doi.org/10.1113/JP278810
3. Alberghina L. The Warburg effect explained: integration of enhanced glycolysis with heterogeneous mitochondria to promote cancer cell proliferation. Int J Mol Sci. 2023;24(21):15787. https://doi.org/10.3390/ijms242115787
4. Liu Y, Sun Y, Guo Y, Shi X, Chen X, Feng W, et al. An overview: the diversified role of mitochondria in cancer metabolism. Int J Biol Sci. 2023;19(3):897–915. https://doi.org/10.7150/ijbs.81609
5. Keoh LQ, Chiu CF, Ramasamy TS. Metabolic plasticity and cancer stem cell metabolism: exploring the glycolysis-OXPHOS switch as a mechanism for resistance and tumorigenesis. Stem Cell Rev Rep. 2025;21(6):2446–2468. https://doi.org/10.1007/s12015-025-10956-y
6. Tang Y, Zhang Z, Chen Y, Qin S, Zhou L, Gao W, et al. Metabolic adaptation-mediated cancer survival and progression in oxidative stress. Antioxidants (Basel). 2022;11(7):1324. https://doi.org/10.3390/antiox11071324
7. Ferreira T, Rodriguez S. Mitochondrial DNA: inherent complexities relevant to genetic analyses. Genes (Basel). 2024;15(5):617. https://doi.org/10.3390/genes15050617
8. Smith ALM, Whitehall JC, Greaves LC. Mitochondrial DNA mutations in ageing and cancer. Mol Oncol. 2022;16(18):3276–3294. https://doi.org/10.1002/1878-0261.13291
9. Gil-de-Gómez L, Balgoma D, Montero O. Lipidomic-based advances in diagnosis and modulation of immune response to cancer. Metabolites. 2020;10(8):332. https://doi.org/10.3390/metabo10080332
10. Boussadia Z, Gambardella AR, Mattei F, Parolini I. Acidic and hypoxic microenvironment in melanoma: impact of tumour exosomes on disease progression. Cells. 2021;10(12):3311. https://doi.org/10.3390/cells10123311
11. Page MJ, McKenzie JE, Bossuyt PM, Boutron I, Hoffmann TC, Mulrow CD, et al. The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. BMJ 2021;372(8284): n71. https://doi.org/10.1136/bmj.n71
12. Stang A. Critical evaluation of the Newcastle-Ottawa scale for the assessment of the quality of nonrandomized studies in meta-analyses. Eur J Epidemiol. 2010;25(9):603–605. https://doi.org/10.1007/s10654-010-9491-z
13. Hooijmans CR, Rovers MM, de Vries RB, Leenaars M, Ritskes-Hoitinga M, Langendam MW. SYRCLE's risk of bias tool for animal studies. BMC Med Res Methodol. 2014;14(1):43. https://doi.org/10.1186/1471-2288-14-43
14. Hu Y, Xu W, Zeng H, He, Lu, Zuo et al. OXPHOS-dependent metabolic reprogramming prompts metastatic potential of breast cancer cells under osteogenic differentiation. Br J Cancer. 2020;123(11):1644–1655. https://doi.org/10.1038/s41416-020-01040-y
15. Evans KW, Yuca E, Scott SS, Zhao M, Paez Arango N, Cruz Pico CX, et al. Oxidative phosphorylation is a metabolic vulnerability in chemotherapy-resistant triple-negative breast cancer. Cancer Res. 2021;81(21):5572–5581. https://doi.org/10.1158/0008-5472.CAN-20-3242
16. El-Botty R, Morriset L, Montaudon E, Tariq, Schnitzler, Bacci et al. Oxidative phosphorylation is a metabolic vulnerability of endocrine therapy and palbociclib resistant metastatic breast cancers. Nat Commun. 2023;14(1):4221. https://doi.org/10.1038/s41467-023-40022-5
17. Vikramdeo KS, Anand S, Sudan SK, Pramanik P, Singh S, Godwin AK, et al. Profiling mitochondrial DNA mutations in tumors and circulating extracellular vesicles of triple-negative breast cancer patients for potential biomarker development. FASEB Bioadv. 2023;5(10):412–426. https://doi.org/10.1096/fba.2023-00070
18. Domínguez-de-la-Cruz E, Muñoz ML, Pérez-Muñoz A, García-Hernández N, Moctezuma-Meza C, Hinojosa-Cruz JC. Reduced mitochondrial DNA copy number is associated with haplogroup and clinical features of breast cancer in Mexican patients. Gene. 2020; 761(1):145047. https://doi.org/10.1016/j.gene.2020.145047
19. Si L, Fu J, Liu W, Hayashi T, Nie Y, Mizuno K, et al. Silibinin inhibits migration and invasion of breast cancer MDA-MB-231 cells through induction of mitochondrial fusion. Mol Cell Biochem. 2020;463(3):189–201. https://doi.org/10.1007/s11010-019-03640-6
20. Martinez-Bernabe T, Pons DG, Oliver J, Sastre-Serra J. Oxidative phosphorylation as a predictive biomarker of oxaliplatin response in colorectal cancer. Biomolecules. 2024;14(11):1359. https://doi.org/10.3390/biom14111359
21. Wu Z, Zuo M, Zeng L, Cui, Liu, Yan et al. OMA1 reprograms metabolism under hypoxia to promote colorectal cancer development. EMBO Rep. 2021;22(12): e50827. https://doi.org/10.15252/embr.202050827
22. Wang C, Cui G, Wang D, Wang M, Chen Q, Wang Y, et al. Crosstalk of oxidative phosphorylation-related subtypes, establishment of a prognostic signature and immune infiltration characteristics in colorectal adenocarcinoma. Cancers (Basel). 2022;14(18):4503. https://doi.org/10.3390/cancers14184503
23. Guo W, Liu Y, Ji X, Guo S, Xie F, Chen Y, et al. Mutational signature of mtDNA confers mechanistic insight into oxidative metabolism remodeling in colorectal cancer. Theranostics. 2023;13(1):324–338. https://doi.org/10.7150/thno.78718
24. Gadicherla R, Rai N, Othayoth R, Kamma S. Characterization of mitochondrial DNA mutations in colorectal cancer progression by in silico approach and use as potential biomarkers for diagnosis and prognosis. Egypt J Med Hum Genet. 2024;25(1):126. https://doi.org/10.1186/s43042-024-00599-y
25. Chen M, Deng S, Cao Y, Wang, Zou, Gu et al. Mitochondrial DNA copy number as a biomarker for guiding adjuvant chemotherapy in stages II and III colorectal cancer patients with mismatch repair deficiency. Ann Surg Oncol. 2024;31(7):6320–6330. https://doi.org/10.1245/s10434-024-15759-y
26. Kobayashi H, Imanaka S. Understanding the impact of mitochondrial DNA mutations on aging and carcinogenesis (review). Int J Mol Med. 2025;56(2):118. https://doi.org/10.3892/ijmm.2025.5559
27. Mei J, Ding P, Gao C, et al. Mitochondrial diseases: molecular pathogenesis and therapeutic advances. MedComm. 2025;6(9):e70385. https://doi.org/10.1002/mco2.70385
28. Du H, Xu T, Yu S, Wu S, Zhang J. Mitochondrial metabolism and cancer therapeutic innovation. Signal Transduct Target Ther. 2025;10(1):245. https://doi.org/10.1038/s41392-025-02311-x
29. Chen K, Wang B, Shu H, Lyu J, Cui W, Fang H. Oxidative phosphorylation at the crossroads of cancer: metabolic orchestration, stromal collusion, and emerging therapeutic horizons. Interdiscip Med. 2025;3(6):e70054. https://doi.org/10.1002/inmd.70054
30. Evans KW, Yuca E, Scott SS, Zhao M, Paez Arango N, Cruz Pico CX, et al. Oxidative phosphorylation is a metabolic vulnerability in chemotherapy-resistant triple-negative breast cancer. Cancer Res. 2021;81(21):5572–5581. https://doi.org/10.1158/0008-5472.CAN-20-3242
31. Guo S, Addison CL. Metabolic vulnerabilities as a therapeutic target in breast cancer. Curr Oncol. 2026;33(2):129. https://doi.org/10.3390/curroncol33020129
32. Chelakkot C, Chelakkot VS, Shin Y, Song K. Modulating glycolysis to improve cancer therapy. Int J Mol Sci. 2023;24(3):2606. https://doi.org/10.3390/ijms24032606
33. Filipp FV, Ratnikov B, De Ingeniis J, Smith JW, Osterman AL, Scott DA. Glutamine-fueled mitochondrial metabolism is decoupled from glycolysis in melanoma. Pigment Cell Melanoma Res. 2012;25(6):732–739. https://doi.org/10.1111/pcmr.12000
34. Machado ND, Heather LC, Harris AL, Higgins GS. Targeting mitochondrial oxidative phosphorylation: lessons, advantages, and opportunities. Br J Cancer. 2023;129(6):897–899. https://doi.org/10.1038/s41416-023-02394-9
35. Li S, Gong J, Kang B, Wang Z, Ma Y, Xia X, et al. Targeting glycolytic metabolism in cancer therapy: current approaches and future perspectives. Cells. 2026;15(4):362. https://doi.org/10.3390/cells15040362
36. Zhao Z, Mei Y, Wang Z, He W. The effect of oxidative phosphorylation on cancer drug resistance. Cancers (Basel). 2022;15(1):62. https://doi.org/10.3390/cancers15010062
37. Tilokani L, Nagashima S, Paupe V, Prudent J. Mitochondrial dynamics: overview of molecular mechanisms. Essays Biochem. 2018;62(3):341–360. https://doi.org/10.1042/EBC20170104
38. Bonora M, Missiroli S, Perrone M, Fiorica F, Pinton P, Giorgi C. Mitochondrial control of genomic instability in cancer. Cancers (Basel). 2021;13(8):1914. https://doi.org/10.3390/cancers13081914
39. Ferreira T, Rodriguez S. Mitochondrial DNA: inherent complexities relevant to genetic analyses. Genes (Basel). 2024;15(5):617. https://doi.org/10.3390/genes15050617
40. Kim HR, Won SJ, Fabian C, Kang MG, Szardenings M, Shin MG. Mitochondrial DNA aberrations and pathophysiological implications in hematopoietic diseases, chronic inflammatory diseases, and cancers. Ann Lab Med. 2015;35(1):1–14. https://doi.org/10.3343/alm.2015.35.1.1